Benchmark¶
Reproducible build / solve / memory comparison between polar-high, linopy, and Pyomo on indexed linear programs, all using HiGHS as the solver. The dense-LP test follows linopy's benchmark, restricted to the linear case so polar-high can run it directly.
The benchmark is a manual artefact, not a CI workflow; runner
variability would make CI numbers misleading. The model code, runner,
and plot script live under
benchmark/
so anyone can re-run on their own hardware in a few minutes.
Headline — dense LP, build-only, single-thread¶
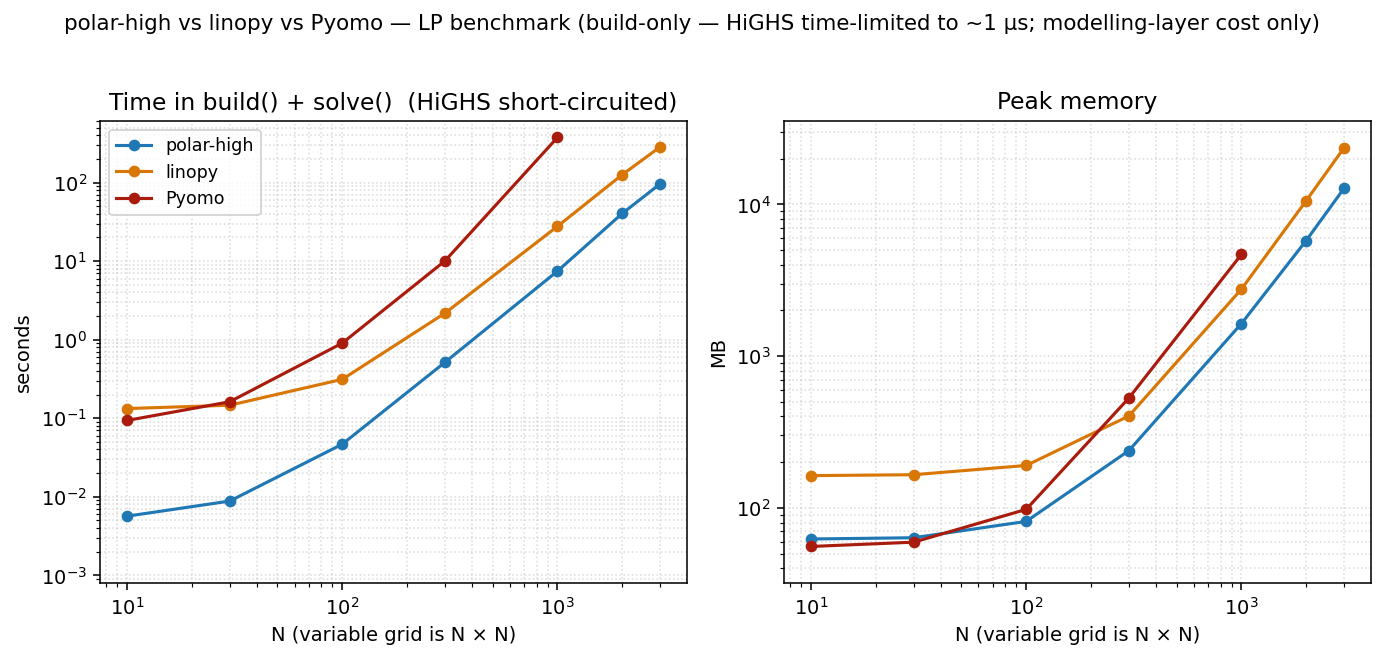
Two panels: total time = build() + solve(), and peak memory.
HiGHS is time-limited to 1 µs so the column reflects pure
modelling-layer cost: what each tool spends turning Python into
something HiGHS can chew on. polar-high and linopy run at
POLARS_MAX_THREADS=1 / OMP_NUM_THREADS=1. Pyomo is
single-threaded by design.
The dense LP that drives this figure (linopy's original model, restricted to LP):
min Σ_{i,j} (2·x[i,j] + y[i,j])
s.t. x[i,j] - y[i,j] >= i for i,j ∈ {1,…,N}
x[i,j] + y[i,j] >= 0 for i,j ∈ {1,…,N}
x, y >= 0
Variables: 2·N². Constraints: 2·N². Closed-form optimum
obj = N²·(N+1), used as the cross-tool correctness anchor.
At N=3000 (a 18 M-variable LP):
| total time | peak memory | |
|---|---|---|
| polar-high | 96 s | 12.7 GB |
| linopy | 290 s | 23.6 GB |
| Pyomo | (caps at N=1000) | (4.7 GB at N=1000) |
polar-high is roughly 3× faster than linopy and uses about half the memory at N=3000. Pyomo's per-coefficient Python overhead makes it unable to keep up past N≈1000 in our 10-minute timeout.
Note on the memory column¶
Peak RSS scales roughly linearly with nnz for all three tools, but
the slope between Pyomo and the columnar tools differs by an order of
magnitude. Pyomo carries each LP coefficient as a small Python
expression-tree node (a VarData reference plus a numeric multiplier
inside a MonomialTermExpression). CPython stores each of those at
roughly 100–200 bytes once header, type pointer, refcount, and per-
object dict are accounted for. polar-high and linopy hold coefficients
as contiguous Arrow / numpy buffers of about 16 bytes per nonzero. At
4 M nonzeros the representation difference compounds to a multi-GB
peak gap.
Between the two columnar tools the comparison flips depending on the test:
- At full HiGHS solve (the linopy-format replication figure
below): linopy's peak at N = 1000 is below polar-high's. xarray's
dense-broadcast representation is genuinely compact on this regular
N × Ngrid, and HiGHS's own working set then dominates everyone's total. - In the build-only headline above: polar-high sits below linopy
at large N. Two factors contribute, in unknown proportion. First,
streaming constraint assembly (polar-high emits LP rows per family
and releases each family's intermediates before the next; linopy
materialises all coefficients at once). Second, the LP-file vs
direct handoff (the build-only test exercises linopy's
io_api="lp"path, the configuration where ourtime_limitshort-circuit reliably reaches HiGHS, which serializes the LP through an intermediate text format; polar-high skips that step viahighspy).
The take-away: which tool wins on memory depends on which phase of the build/solve cycle dominates your run. Re-run on your own model before assuming any of these ratios apply.
Reproducing¶
git clone https://github.com/nodal-tools/polar-high.git
cd polar-high
pip install -r benchmark/requirements.txt
python benchmark/run.py # ~10 min on a laptop
python benchmark/plot.py # writes docs/assets/benchmark*.png
benchmark/run.py accepts --sizes, --tools, --threads,
--repeats, --time-limit. Each (tool, N) cell runs in a fresh
subprocess so caches and Python state don't carry over. The plot
takes the median across reps. See
benchmark/README.md
for the methodology details.
Threading: defaults and where it matters¶
polar-high sets POLARS_MAX_THREADS=1 at import. The data below
shows why.
Scaling with threads at small N¶
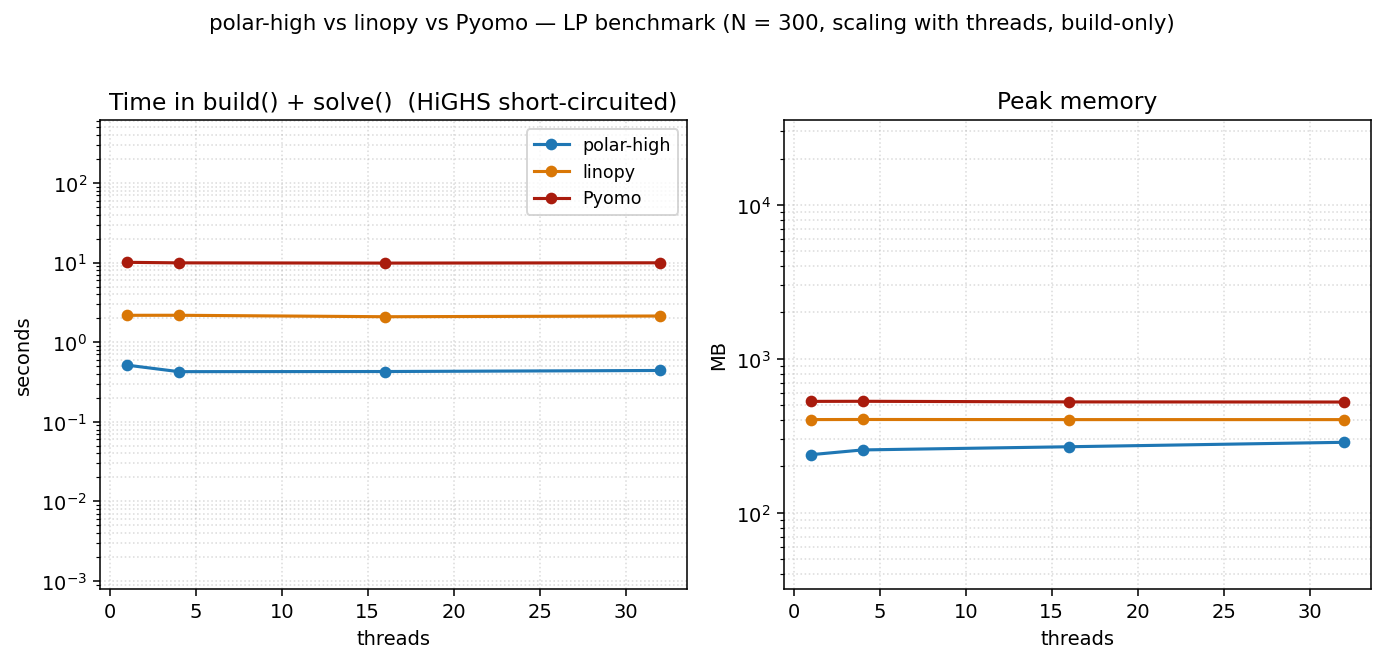
At a small problem size (N=300) the build is too small to amortize Rayon's coordination overhead. All three tools are essentially flat across thread counts. polars's parallelism doesn't pay; linopy and Pyomo are single-threaded by design. polar-high's peak memory rises slightly with threads (per-thread scratch buffers, ~1.5 MB per thread on this LP) without a corresponding time win.
Scaling with threads on a larger LP¶
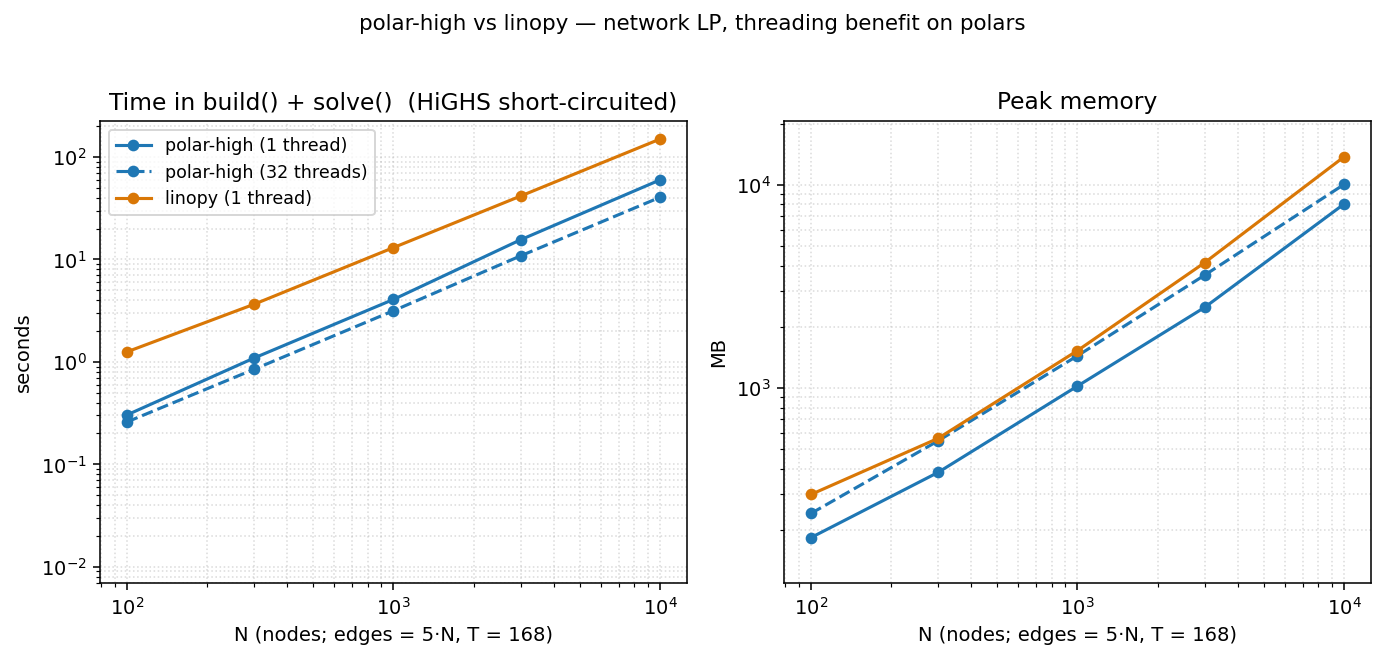
On the network LP (irregular topology, T=168, see below), polar's threading does start paying as N grows, and grows steadily:
| N | polar 1 thread | polar 32 threads | speedup |
|---|---|---|---|
| 100 | 0.30 s | 0.26 s | 1.16× |
| 300 | 1.08 s | 0.84 s | 1.29× |
| 1 000 | 4.08 s | 3.18 s | 1.28× |
| 3 000 | 15.5 s | 10.8 s | 1.43× |
| 10 000 | 60.0 s | 40.5 s | 1.48× |
At threads=32, polar uses ~25 % more memory than at threads=1 (per-thread Rayon scratch). The takeaway:
- Default threads=1 is the right choice for typical workloads. Faster and leaner on small/medium LPs; only modestly slower than threads=32 even at large N.
- Override to threads=N by setting
POLARS_MAX_THREADSbefore importing polar-high. Worth doing for genuinely large indexed LPs where you've checked that the polars-side build is the bottleneck.
Network LP: irregular topology¶
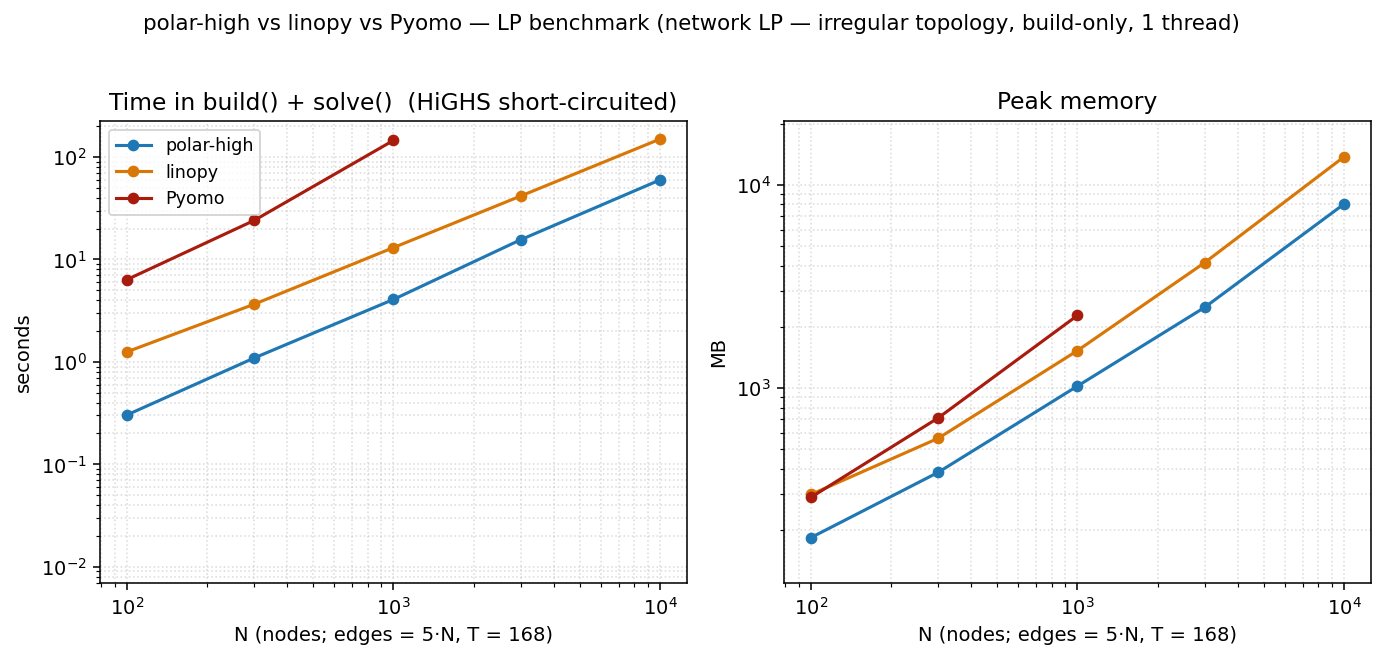
The dense N × N headline above is the model linopy's benchmark
uses, and it's flattering to xarray: every cell has the same shape,
indexing is rectangular, and dense numpy arrays compress the
coefficient matrix. Real indexed LPs (energy systems, supply chains)
aren't shaped that way. They have irregular topology: flow
variables on edges, balance constraints on nodes, with an edge→node
lookup that varies per node.
The network LP captures that:
N nodes, E = 5·N edges drawn at random (Hamiltonian cycle + 4·N random),
T = 168 time steps (one week of hours)
vars flow[e, t] >= 0
cstr 1 flow[e, t] <= cap[e, t] ∀ (e, t)
cstr 2 Σ_{e: dst[e]=n} flow[e,t] − Σ_{e: src[e]=n} flow[e,t]
== demand[n, t] ∀ (n, t)
obj min Σ cost[e] · flow[e, t]
The node-balance constraint requires an edge→node lookup. polar-high
expresses this as a polars join (Where(flow, edges_dst_n)); linopy
uses flow.groupby(dst_array).sum(); Pyomo precomputes
incoming[n] adjacency dicts and iterates them. Each idiom is
natural to its tool.
At N=10 000 (≈ 8.4 M variables, ≈ 10 M constraints):
| total time | peak memory | |
|---|---|---|
| polar-high | 60 s | 8.0 GB |
| linopy | 146 s | 13.7 GB |
| Pyomo | timed out at N=3000 | — |
polar wins all three columns and Pyomo can't finish at this scale
inside our 10-minute timeout. Notably, linopy's memory advantage on
the dense LP doesn't carry over here. The irregular (e, t) →
(n, t) join breaks xarray's dense-broadcast pattern, while polar's
join semantics map naturally onto the operation.
Replication: linopy's original benchmark format¶
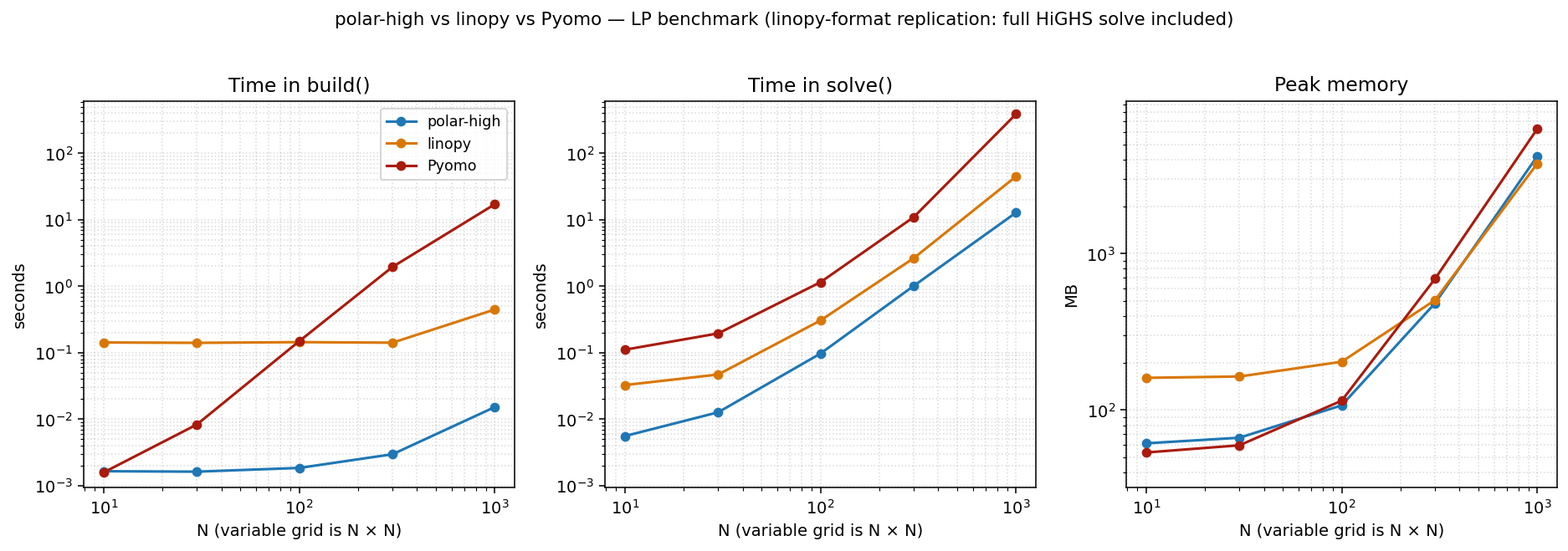
This is the same dense LP at threads=1 in the linopy benchmark's original three-panel format: build / solve / memory shown separately, with the full HiGHS solve included (no time limit). We include it for direct visual comparison with linopy's published numbers and as the methodology footnote.
Caveat: read with care. The build() / solve() split is not
an apples-to-apples decomposition across tools. Each library draws
the boundary in a different place:
- polar-high's
build()registers metadata only. The matrix assembly happens insidesolve()alongside the HiGHS call. - linopy's
build()materialises xarray coefficient arrays eagerly;solve()writes them to an LP file and hands it to HiGHS. - Pyomo's
build()instantiatesVarData/ConstraintData/MonomialTermExpressionPython objects per coefficient. Most of the work happens here;solve()walks the expression tree to emit the LP.
So when you compare the build column between tools, you're
comparing what each tool chose to put in build(). The
apples-to-apples cross-tool measurement is build() + solve(),
which is the headline figure at the top of this page.
That said, the replication is faithful: the closed-form objective is hit exactly, the model is the same LP, and HiGHS solves the same sparse matrix. linopy's numbers reproduce within run-to-run noise.
Fairness caveats¶
- Each tool builds the LP in its own natural style; the closed-form objective (or cross-tool agreement on the network LP) confirms they're all solving the same problem.
- Pyomo's solve uses the
appsi_highspersistent-solver interface, the fastest Pyomo→HiGHS path. The file-basedSolverFactory("highs")route would inflate Pyomo's solve time with LP-write/reread overhead. - linopy uses
io_api="lp"explicitly: the configuration where ourtime_limitshort-circuit reliably reaches HiGHS for the build-only test. linopy'sio_api="direct"is faster on full solves but interacts awkwardly with the time-limit short-circuit. - polar-high's solve calls into
highspydirectly with no intermediate file in either configuration. - HiGHS itself stays at 1 thread across all tools, so the solve column remains apples-to-apples.
- Each
(tool, N)cell runs in a fresh Python process, so caches and memory don't carry over between cells. Python startup costs ~50 ms per cell, negligible at our sizes. - One machine, one Python version, one HiGHS version. Re-run on your own hardware before reading too much into the absolute numbers; the ratios between tools are what's portable.
What's not in the benchmark¶
- MIP, QP, warm starts, and Lagrangian decomposition. The benchmark deliberately exercises the matrix-assembly path on plain LPs.
- JuMP and gurobipy. Both have toolchain or licence friction
that defeats "anyone can re-run from a clean
pip install".
If you'd like a different model class (block-angular, time-coupled, mixed-integer with branching) in the benchmark, open an issue. The runner is a small enough surface to extend.